This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
DNA Motifs
MOTIFBy using MOTIF program, eleven DNA motifs could be found for the full-length HEXA gene. The following list shows the summary of the 11 DNA motifs.
|
|
Motif: CTCK_1
Prosite ID: PS01185 Description: C-terminal cystine knot signature. Pattern: C-C-x(13)-C-x(2)-[GN]-x(12)-C-x-C-x(2,4)-C. |
Motif: ANAPHYLATOXIN_1
Prosite ID: PS01177 Description: Anaphylatoxin domain signature. Pattern: [CSH]-Cx(2)-[GAP]-x(7,8)-[GASTDEQR]-C-[GASTDEQL]- x(3,9)-[GASTDEQN]-x(2)-[CE]-x(6,7)-C-C. |
|
Motif: IGFBP_N_1
Prosite ID: PS00222 Description: Insulin-like growth factor-binding protein (IGFBP) N-terminal domain signature. Pattern: [GP]-C-[GSET]-[CE]-[CA]-x(2)-C-[ALP]-x(6)-C. |
Motif: THIOLASE_3
Prosite ID: PS00099 Description: Thiolases active site. Pattern: [AG]-[LIVMA]-[STAGCLIVM]-[STAG]-[LIVMA]-C-{Q}-[AG]-x- [AG]-x-[AG]-x-[SAG]. |
|
Motif: 4FE4S_FER_1
PrositeID: PS00198 Description: 4Fe-4S ferredoxin-type iron-sulfur binding region signature. Pattern: C-x-{P}-C-{C}-x-C-{CP}-x-{C}-C-[PEG]. |
Motif: 2FE2S_FER_1
PrositeID: PS00197 Description: 2Fe-2S ferredoxin-type iron-sulfur binding region signature. Pattern: C-{C}-{C}-[GA]-{C}-C-[GAST]-{CPDEKRHFYW}-C. |
Motif: DEFENSIN
PrositeID: PS00269
Description: Mammalian defensins signature.
Pattern: C-x-C-x(3,5)-C-x(7)-G-x-C-x(9)-C-C.
PrositeID: PS00269
Description: Mammalian defensins signature.
Pattern: C-x-C-x(3,5)-C-x(7)-G-x-C-x(9)-C-C.
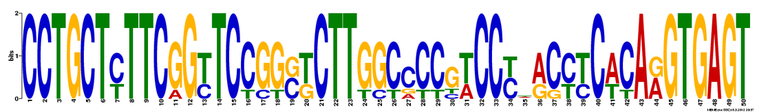
MEME
60,000 bp maximum did not allow searching the gene sequence. Thus, I was not able to input all the gene sequences of HEXA gene homologs into MEME. Instead of using gene sequence, I inputed HEXA protein sequences of all homologs into MEME database. Three motifs were found and they were shown in the following figures. Click here to look at results from MEME.
|
Analysis
I used two programs available for determining DNA motifs: MOTIF and MEME. MEME is helpful in locating potential motifs but does not appear to be as helpful as MOTIF Search in identifying them. However, I still could not find significant description and information about the importance of protein interactions for a particular region .
References
1. MOTIF
2. MEME
3. Bailey, T. & Elkan, C. (1994) "Fitting a mixture model by expectation maximization to discover motifs in biopolymers", Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, AAAI Press, Menlo Park, California, 28-36. [PUBMED]
1. MOTIF
2. MEME
3. Bailey, T. & Elkan, C. (1994) "Fitting a mixture model by expectation maximization to discover motifs in biopolymers", Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, AAAI Press, Menlo Park, California, 28-36. [PUBMED]