This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
HEXA Protein Domains
|
Pfam
Searching on Pfam by using the full sequence of HEXA protein resulted in a map of the protein shown below. By setting Pfam-A E-value cut-off to 1.0, Pfam found 2 Pfam-A matches to the HEXA protein sequence and these 2 matches are significant. |
Functions of each domain:
The green domain represents glycoside hydrolase family 20, domain 2, with an e-value of 4.2e-43. Pfam suggested that the location of this domain to be from 35 to 165. This domain has a zincin-like fold [2]. Beta-hexosaminidase is one of the known enzymes in Glycoside hydrolase family 20 CAZY. In the brain and other tissues, beta-hexosaminidase A degrades GM2 gangliosides [3].
The red domain represents glycosyl hydrolase family 20, catalytic domain, with an e-value of 2e-84. This domain located between 167 and 487, which has a TIM barrel fold [2].
The purple diamond in the figure also indicates the Pfam predicted active site which is the residue E323. This pedicted active site is located at 323.
The green domain represents glycoside hydrolase family 20, domain 2, with an e-value of 4.2e-43. Pfam suggested that the location of this domain to be from 35 to 165. This domain has a zincin-like fold [2]. Beta-hexosaminidase is one of the known enzymes in Glycoside hydrolase family 20 CAZY. In the brain and other tissues, beta-hexosaminidase A degrades GM2 gangliosides [3].
The red domain represents glycosyl hydrolase family 20, catalytic domain, with an e-value of 2e-84. This domain located between 167 and 487, which has a TIM barrel fold [2].
The purple diamond in the figure also indicates the Pfam predicted active site which is the residue E323. This pedicted active site is located at 323.
SMART

SMART was used to analyze the HEXA sequence. Results shown by SMART were slightly different from that of Pfam.
The Glyco_hydro_20b domain within the HEXA protein sequence starts at position 35 and ends at position 165. The E-value for the Glyco_hydro_20b domain shown below is 1.3e-44. The second domain which starts at position 167 and ends at position 488 of the protein sequence is Glyco_hydro_20 ; the E-value for the Glyco_hydro_20 domain shown below is 1.00e-85. Segments of low compositional complexity is determined by the SEG program (in purple). The region of low compositional complexity chosen starts at position 4 of the query sequence and ends at position 24. Intron positions are indicated with vertical lines showing the intron phase and exact position in AA.
The Glyco_hydro_20b domain within the HEXA protein sequence starts at position 35 and ends at position 165. The E-value for the Glyco_hydro_20b domain shown below is 1.3e-44. The second domain which starts at position 167 and ends at position 488 of the protein sequence is Glyco_hydro_20 ; the E-value for the Glyco_hydro_20 domain shown below is 1.00e-85. Segments of low compositional complexity is determined by the SEG program (in purple). The region of low compositional complexity chosen starts at position 4 of the query sequence and ends at position 24. Intron positions are indicated with vertical lines showing the intron phase and exact position in AA.
Prosite

By using Prosite, I entered the HEXA protein sequence and unchecked "exclude patterns with a high probability of occurrence". The Prosite utilized "hits by profiles with a high probability of occurrence" to find motifs on HEXA sequence. However, Prosite only stated that there were 23 hits (by 7 distinct patterns) on 1 sequence. It did not identify significant location for particular domain.
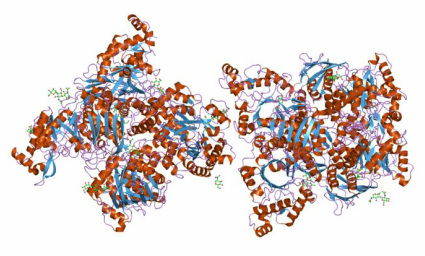
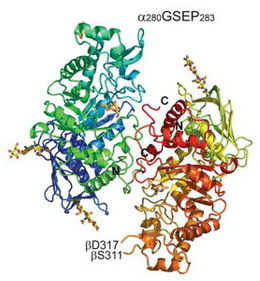
Structure of Glycosyl hydrolase family 20 Domain of HEXA
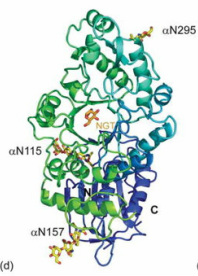
Hex A is a member of the Family 20 glycoside hydrolases (glycosidase). The α-subunit and β-subunit of human Hex A are encoded by the related genes HEXA and HEXB, respectively. However, the hydrolysis of GM2 ganglioside is catalyzed only by the α-subunit of Hex A. There are three glycosylation sites on the α-subunit of Hex A which are αAsn115, αAsn157 and αAsn295. Lemieux M. et al. point out that α-subunit has a lower inherent stability, which make the α-subunit more susceptible to the missense mutations. This hypothesis is supported by the In vitro mutagenesis and expression experiments that duplicated α-point mutation in the aligned site with the β-subunit. Those residues involved in chronic TSD are mostly located at domain 2 of the α-subunit.
Analysis
Based on the Pfam and SMART database, two domains are found on the HEXA sequence. They are glycoside hydrolase family 20, domain 2 and glycosyl hydrolase family 20, catalytic domain. The E-value given by SMART is lower than the one provided by Pfam. Although the names of the domains given by these two database were slightly different, Pfam and SMART identified the almost same location for the domains.
According to Lemieux, M. et al., the majority of residues involved in acute TSD are located throughout domain II of the α-subunit, distributed along the β-strands and helices comprising the TIM barrel [6]. The B1 variant mutation occurs mainly at αArg178 [7]. The hydrogen bonding network in the active site is disrupted. It can reduce GM2 binding and severely affect the activity of the mutant α-active site of Hex A. Patients with homozygous αArg178His have a sub-acute phenotype of TSD, whereas heterozygotes with a second null allele shows more severe acute phenotype. Furthermore, the αAsp258His mutation was identified in a TSD patient with a severe subacute phenotype [8]. The αGly269Ser is the predominant mutation found in adult TSD (ATSD).
At the end of the paper written by Lemieux, M. et al., the authors points out that N-acetylglucosamine-thiazoline (NGT), a mechanism-based inhibitor bound to Hex A, can act as a chemical chaperone. In this case, it can prevent misfolding of a Hex A mutant associated with adult onset TSD. As a result, the residual activity of Hex A can be catalyzed and hence preventing TSD.
According to Lemieux, M. et al., the majority of residues involved in acute TSD are located throughout domain II of the α-subunit, distributed along the β-strands and helices comprising the TIM barrel [6]. The B1 variant mutation occurs mainly at αArg178 [7]. The hydrogen bonding network in the active site is disrupted. It can reduce GM2 binding and severely affect the activity of the mutant α-active site of Hex A. Patients with homozygous αArg178His have a sub-acute phenotype of TSD, whereas heterozygotes with a second null allele shows more severe acute phenotype. Furthermore, the αAsp258His mutation was identified in a TSD patient with a severe subacute phenotype [8]. The αGly269Ser is the predominant mutation found in adult TSD (ATSD).
At the end of the paper written by Lemieux, M. et al., the authors points out that N-acetylglucosamine-thiazoline (NGT), a mechanism-based inhibitor bound to Hex A, can act as a chemical chaperone. In this case, it can prevent misfolding of a Hex A mutant associated with adult onset TSD. As a result, the residual activity of Hex A can be catalyzed and hence preventing TSD.
References
1. Pfam
2. Tews, I. , Perrakis, A. , Oppenheim, A. , Dauter, Z. , Wilson, K. , et al. (1996). Bacterial chitobiase structure provides insight into catalytic mechanism and the basis of tay-sachs disease. Nat Struct Biol, 3(7), 638-648. PUBMED:8673609
3. Interpro database
4. SMART
5. Prosite
6. Lemieux, M. , Mark, B. , Cherney, M. , Withers, S. , Mahuran, D. , et al. (2006). Crystallographic structure of human beta-hexosaminidase a: Interpretation of tay-sachs mutations and loss of gm2 ganglioside hydrolysis. J Mol Biol, 359(4), 913-929. doi: 10.1016/j.jmb.2006.04.004
7. Ohno K, Suzuki K. Mutation in GM2-gangliosidosis B1 variant. J Neurochem. 1988;50:316–318. [PubMed]
8. Bayleran J, Hechtman P, Kolodny E, Kaback M. Tay-Sachs disease with hexosaminidase A: characterization of the defective enzyme in two patients. Am J Hum Genet. 1987;41:532–548. [PubMed]
2. Tews, I. , Perrakis, A. , Oppenheim, A. , Dauter, Z. , Wilson, K. , et al. (1996). Bacterial chitobiase structure provides insight into catalytic mechanism and the basis of tay-sachs disease. Nat Struct Biol, 3(7), 638-648. PUBMED:8673609
3. Interpro database
4. SMART
5. Prosite
6. Lemieux, M. , Mark, B. , Cherney, M. , Withers, S. , Mahuran, D. , et al. (2006). Crystallographic structure of human beta-hexosaminidase a: Interpretation of tay-sachs mutations and loss of gm2 ganglioside hydrolysis. J Mol Biol, 359(4), 913-929. doi: 10.1016/j.jmb.2006.04.004
7. Ohno K, Suzuki K. Mutation in GM2-gangliosidosis B1 variant. J Neurochem. 1988;50:316–318. [PubMed]
8. Bayleran J, Hechtman P, Kolodny E, Kaback M. Tay-Sachs disease with hexosaminidase A: characterization of the defective enzyme in two patients. Am J Hum Genet. 1987;41:532–548. [PubMed]