This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Organisms Phenotypes and RNAi
The RNAi provides a great opportunity in the basic biological research. Its pathway is exploited in experimental biology to study the function of genes in in vitro (cell culture) and in vivo in model organisms. The Tay-Sachs disease is resulted from the mutation in human HEXA gene. Studies have been done on model organisms to study the relationship between the HEXA mutant and RNAi phenotypes in humans versus model organisms. RNA interference (RNAi) is performed to knock out the homologous HEXA gene in each organism. The model organisms with phenotypic similarity to the TSD associated to human HEXA can help researchers in studying how the mutant gene affect this disorder.
The RNAi phenotypes from model organisms are shown in the following section:
Mus musculus (mouse)
Targeted (knockout) Allele: Hexa<tm1Rlp>
In Yamanaka S. et al. article, they introduced the mice in which totally deficient in beta-hexosaminidase A through disruption of the mouse HEXA gene by homologous recombination.
Targeted (knockout) Allele: Hexa<tm1Rlp>
- Abnormal brain morphology [3]
- Abnormal neuron morphology [3]
In Yamanaka S. et al. article, they introduced the mice in which totally deficient in beta-hexosaminidase A through disruption of the mouse HEXA gene by homologous recombination.
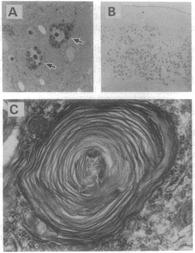
Figure 1: Hexa -/- mice show the neuropathology characteristic of Tay-Sachs disease. (A)membranous cytoplasmic bodies (MCBs) are clearly recognized in the perikarya of two neurons in the parietal cortex from a Hexa -/- mouse. A 1-lm E (B) Many neurons are immunostained with anti-Gm2 ganglioside antibody in a 7-pam cryosection of the mamillary body from an
8-month-old mouse. Note the dark peroxidase reaction product staining the neurons. Sections from control mice were totally negative for the reaction product. (x30.) (C) Electron micrograph of an
MCB composed of multilayered lamellae in a cerebral cortical neuron from a 7-week-old Hexa -/- mouse. (x9500.)
Figures obtained from Yamanaka S. et al.
|
Figures obtained from Jeyakumar, M. et al.
|
Danio rerio (zebrafish)
ZFIN did not show any result for RNAi experiment done on zebra fish. No data is available for its phenotype, mutants and targeted knockdown. Click on hexosaminidase A (alpha polypeptide) for the search result.
ZFIN did not show any result for RNAi experiment done on zebra fish. No data is available for its phenotype, mutants and targeted knockdown. Click on hexosaminidase A (alpha polypeptide) for the search result.
Caenorhabditis elegans (Worm)
The Blast Search in WormBase was used to find the gene homolog of HEXA by entering the protein sequences. However, the following targeted gene is the top hit that I received from this search but the study was not related to the Tay-Sachs disease.
Targeted gene: hex-1
Description:hex-1 encodes a beta-N-acetylhexosaminidase that is orthologous to the human gene CERVICAL CANCER PROTO-ONCOGENE 7 (HEXB; OMIM:606873), which when mutated leads to disease. [Gutternigg M et al., 2007]
However, no abnormalities were observed via RNAi of hex-1. [Kamath RS et al.]
The Blast Search in WormBase was used to find the gene homolog of HEXA by entering the protein sequences. However, the following targeted gene is the top hit that I received from this search but the study was not related to the Tay-Sachs disease.
Targeted gene: hex-1
Description:hex-1 encodes a beta-N-acetylhexosaminidase that is orthologous to the human gene CERVICAL CANCER PROTO-ONCOGENE 7 (HEXB; OMIM:606873), which when mutated leads to disease. [Gutternigg M et al., 2007]
However, no abnormalities were observed via RNAi of hex-1. [Kamath RS et al.]
Drosophila Melanogaster (Fruit Fly)
FlyBase was used to identify the phenotypes caused by the knock out of the homologous HEXA gene. However, no RNAi information related to TSD for D. melanogaster . The follwing result is the top hit I received from Flybase. Click on the gene name for information.
Targeted gene: Dmel\Hex-A
Phenotype : electrophoretic variant
FlyBase was used to identify the phenotypes caused by the knock out of the homologous HEXA gene. However, no RNAi information related to TSD for D. melanogaster . The follwing result is the top hit I received from Flybase. Click on the gene name for information.
Targeted gene: Dmel\Hex-A
Phenotype : electrophoretic variant
Analysis
According to the analysis on all the model organisms above from each database, the mutant phenotypes expressed in the mice are the closest characteristics of the human Tay-Sachs disease. It might be more difficult to perform RNAi in other organisms. This is perhaps because TSD is not caused by a single type of mutation. Instead, the clinical heterogeneity manifestations of TSD are due to different mutations which affect the catalytic activity of β-hexosaminidase in variable degrees.
According to Jeyakumar, M. et al., the Tay–Sachs (Hexa-/-) mouse model lacks an phenotype as mice can escape from TSD by partially bypass the blocked catabolic pathway. Thus, I think that Mus musculus (mouse) could be the most appropriate model organism that we can choose to perform RNAi experiment.
According to Jeyakumar, M. et al., the Tay–Sachs (Hexa-/-) mouse model lacks an phenotype as mice can escape from TSD by partially bypass the blocked catabolic pathway. Thus, I think that Mus musculus (mouse) could be the most appropriate model organism that we can choose to perform RNAi experiment.
References
1. Kabir, M. , Qadir, S. , Hassan, S. , Ahn, J. , & Wang, M. (4784). Rnai: An emerging field of molecular research. African Journal of Biotechnology, 7(25), 4784-4788. From http://www.ajol.info/index.php/ajb/article/viewFile/59671/47957
2. MGI
3. Yamanaka, S. , Johnson, M. , Grinberg, A. , Westphal, H. , Crawley, J. , et al. (1994). Targeted disruption of the hexa gene results in mice with biochemical and pathologic features of tay-sachs disease. Proceedings of the National Academy of Sciences of the United States of America, 91(21), 9975-9979. [PUBMED]
4. ZFIN
5. Wormbase
6. FlyBase
7. Jeyakumar, M. , Smith, D. , Eliott-Smith, E. , Cortina-Borja, M. , Reinkensmeier, G. , et al. (2002). An inducible mouse model of late onset tay-sachs disease. Neurobiology of Disease, 10(3), 201-210. [PUBMED]
1. Kabir, M. , Qadir, S. , Hassan, S. , Ahn, J. , & Wang, M. (4784). Rnai: An emerging field of molecular research. African Journal of Biotechnology, 7(25), 4784-4788. From http://www.ajol.info/index.php/ajb/article/viewFile/59671/47957
2. MGI
3. Yamanaka, S. , Johnson, M. , Grinberg, A. , Westphal, H. , Crawley, J. , et al. (1994). Targeted disruption of the hexa gene results in mice with biochemical and pathologic features of tay-sachs disease. Proceedings of the National Academy of Sciences of the United States of America, 91(21), 9975-9979. [PUBMED]
4. ZFIN
5. Wormbase
6. FlyBase
7. Jeyakumar, M. , Smith, D. , Eliott-Smith, E. , Cortina-Borja, M. , Reinkensmeier, G. , et al. (2002). An inducible mouse model of late onset tay-sachs disease. Neurobiology of Disease, 10(3), 201-210. [PUBMED]

