This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Below is a copy of my final presentation in both Powerpoint and PDF format.
|
|
||||||||||||
Conclusions
The purpose of this project is to analyze how the HEXA gene affects Tay-Sachs disease (TSD) patients by using several bioinformatics programs and databases found on the Internet. HEXA is conserved mainly in eukaryotes, including human, chimpanzee, dog, cattle, mouse, rat, chicken, zebrafish, and A.thaliana. Organisms that possess HEXA gene homologs can be useful model organisms for research purposes. In Yamanaka S. et al. article, they introduced the mice in which totally deficient in beta-hexosaminidase A through disruption of the mouse HEXA gene by homologous recombination. Mice has been chosen as the model organism because the mutant phenotypes expressed in the mice are the closest characteristics of the human Tay-Sachs disease. For example, membranous cytoplasmic bodies (MCBs) can be observed in the parietal cortex of Hexa -/- mice. MCBs are the ganglioside stored in the form of concentrically arranged lamellae. We can also oberve the loss of motor skill in the mutant mice. Besides, mouse HEXA is 84% homologous to the human copy.
Upon looking at HEXA gene ontology, I discovered that HEXA involve in the ganglioside catabolic process (biological process) and beta-N-acetylhexosaminidase activity (molecular function). Further investigation of HEXA protein domains allowed me to discover glycoside hydrolase family 20, domain 2 and glycosyl hydrolase family 20, catalytic domain. According to Lemieux, M. et al., the majority of residues involved in acute TSD are located throughout domain II of the α-subunit, distributed along the β-strands. The hydrolysis of GM2 ganglioside is catalyzed only by the α-subunit of Hex A.
I decided to further investigate the proteins involved in the degradation of GM2 in the HEXA interaction network.
Experimental question #1: What is the GO for the proteins involved in the degradation of GM2 in the HEXA interaction network?
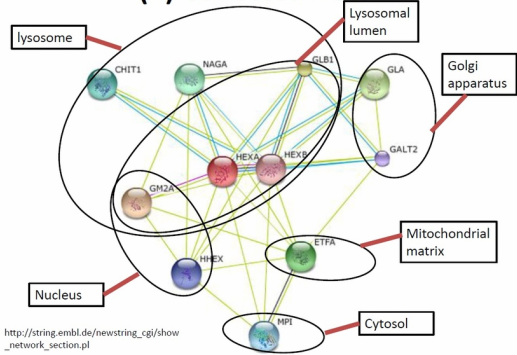
Hypothesis: The GO for the related proteins in the HEXA interaction network is categorized based on cellular component. There are categorized into groups like lysosome, lysosomal lumen, membrane, nucleus, cytosol, Golgi apparatus, and mitochondrial matrix.
Method: Using the AMIGO database to find out the gene ontology of each related proteins in the degradation of GM2 and categorize them into different cellular component.
Result: Most of the proteins in the HEXA interaction network are located at the lysosome and lysosomel lumen.
Upon looking at HEXA gene ontology, I discovered that HEXA involve in the ganglioside catabolic process (biological process) and beta-N-acetylhexosaminidase activity (molecular function). Further investigation of HEXA protein domains allowed me to discover glycoside hydrolase family 20, domain 2 and glycosyl hydrolase family 20, catalytic domain. According to Lemieux, M. et al., the majority of residues involved in acute TSD are located throughout domain II of the α-subunit, distributed along the β-strands. The hydrolysis of GM2 ganglioside is catalyzed only by the α-subunit of Hex A.
I decided to further investigate the proteins involved in the degradation of GM2 in the HEXA interaction network.
Experimental question #1: What is the GO for the proteins involved in the degradation of GM2 in the HEXA interaction network?
Hypothesis: The GO for the related proteins in the HEXA interaction network is categorized based on cellular component. There are categorized into groups like lysosome, lysosomal lumen, membrane, nucleus, cytosol, Golgi apparatus, and mitochondrial matrix.
Method: Using the AMIGO database to find out the gene ontology of each related proteins in the degradation of GM2 and categorize them into different cellular component.
Result: Most of the proteins in the HEXA interaction network are located at the lysosome and lysosomel lumen.
Experimental question #2: What domains are found in the related proteins located at the lysosome? Are Glycoside hydrolase family 20, catalytic domain found in those proteins?
Hypothesis: The Glycoside hydrolase family, catalytic domain should be observed in some of the proteins located at the lysosome in the HEXA interaction network because this domain play an important role in degrading GM2 ganglioside.
Method: Using the SMART database to find the domains of the related proteins that are located in the lysosome.
Resut: The Glycoside hydrolase family, catalytic domain can be observed in the proteins located at the lysosme such as HEXB, CHIT1, and GLB1.
In summary, Tay-Sachs Disease (TSD) is an autosomal recessive disease caused by mutations in both alleles of a gene (HEXA) on chromosome 15. The Glycoside hydrolase family 20, catalytic domain could play an important role in degrading GM2 ganglioside.
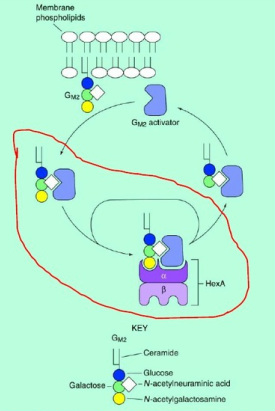
Figure 2: Model for the lysosomal metabolism of GM2: HexA is a heterodimer. It does not interact directly with the membrane-bound GM2. Instead, the GM2 activator protein extracts the glycolipid, and the resulting activator-lipid complex is the substrate for the enzymatic reaction catalyzed by HexA (removal of the terminal N-acetylgalactosamine from GM2).
Future Directions
1. The figure 2 shows the model for the lysosomal metabolism of GM2. I would like to know the mechanism of action of GM2-AP that can cause the recognition of Hex-A (the stage circled with red line in the figure 2).
Possible Method: TAP-tag - Label GM2-AP and determine how it is recognize HEXA
2. Chaperone Therapy
- I would like to use different kind of chaperone to treat TSD with different mutant variations.
3. I would like to create effective GM2 ganglioside inhibitors to treat TSD.
Possible Method: TAP-tag - Label GM2-AP and determine how it is recognize HEXA
2. Chaperone Therapy
- I would like to use different kind of chaperone to treat TSD with different mutant variations.
3. I would like to create effective GM2 ganglioside inhibitors to treat TSD.
References
1. Jeyakumar, M. , Smith, D. , Eliott-Smith, E. , Cortina-Borja, M. , Reinkensmeier, G. , et al. (2002). An inducible mouse model of late onset tay-sachs disease. Neurobiology of Disease,10(3), 201-210.
2. Kabir, M. , Qadir, S. , Hassan, S. , Ahn, J. , & Wang, M. (4784). Rnai: An emerging field of molecular research. African Journal of Biotechnology, 7(25), 4784-4788. From http://www.ajol.info/index.php/ajb/article/viewFile/59671/47957
3. MGI
4. Yamanaka, S. , Johnson, M. , Grinberg, A. , Westphal, H. , Crawley, J. , et al. (1994). Targeted disruption of the hexa gene results in mice with biochemical and pathologic features of tay-sachs disease. Proceedings of the National Academy of Sciences of the United States of America, 91(21), 9975-9979. [PUBMED]
5. National Tay-Sachs & Allied Diseases
6. String: http://string.embl.de
7. SMART: http://smart.embl-heidelberg.de/
8. PFAM: http://pfam.sanger.ac.uk/
2. Kabir, M. , Qadir, S. , Hassan, S. , Ahn, J. , & Wang, M. (4784). Rnai: An emerging field of molecular research. African Journal of Biotechnology, 7(25), 4784-4788. From http://www.ajol.info/index.php/ajb/article/viewFile/59671/47957
3. MGI
4. Yamanaka, S. , Johnson, M. , Grinberg, A. , Westphal, H. , Crawley, J. , et al. (1994). Targeted disruption of the hexa gene results in mice with biochemical and pathologic features of tay-sachs disease. Proceedings of the National Academy of Sciences of the United States of America, 91(21), 9975-9979. [PUBMED]
5. National Tay-Sachs & Allied Diseases
6. String: http://string.embl.de
7. SMART: http://smart.embl-heidelberg.de/
8. PFAM: http://pfam.sanger.ac.uk/